100
Benchmark Coverage
 NeuroClaw
NeuroClaw
NeuroClaw turns neuroimaging workflows into executable, iterative, and reproducible research loops across data preparation, model execution, validation, and refinement.
Benchmark Coverage
base-subagent-interface
Dataset-Aware Orchestration
Execution and Audit Logs
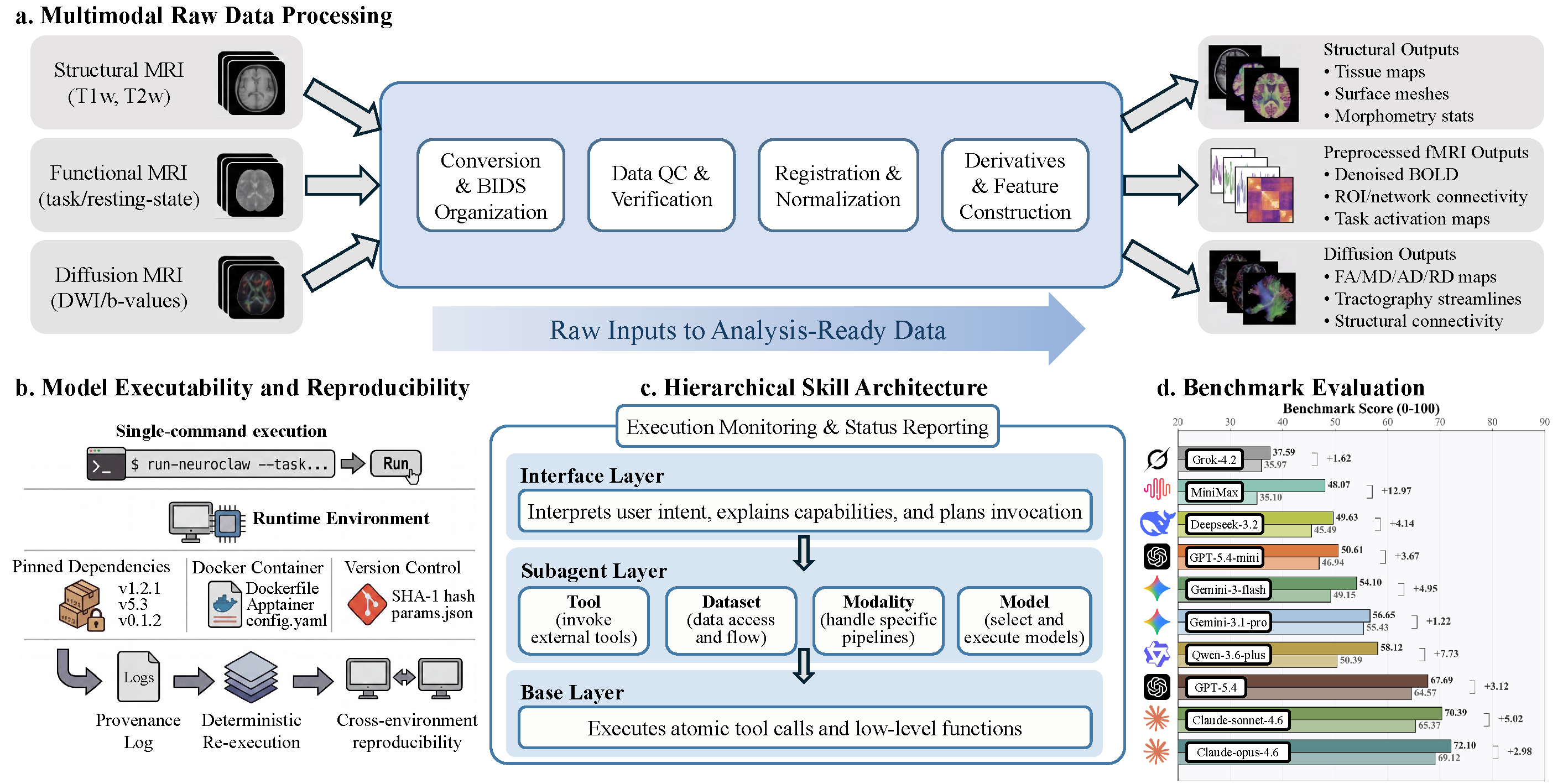
NeuroClaw starts from raw multimodal data and uses dataset semantics, BIDS metadata, and workflow stage context to keep downstream analyses scientifically coherent.
Managed Python environments, Docker support, GPU setup, checkpoints, artifact checks, and audit logs make neuroimaging toolchains easier to rerun and verify.
A three-tier interface, subagent, and base-skill design decomposes long neuroimaging workflows into controlled, reusable operations across tools, modalities, datasets, and models.
NeuroBench evaluates planning completeness, tool-use reasonableness, and command/code correctness under realistic neuroimaging tasks with and without NeuroClaw skills.
1.1 Data I/O and BIDS: BIDS organization, DICOM→NIfTI, NIfTI→DICOM, metadata checks, and low-level NIfTI/FreeSurfer I/O with Nibabel.
1.2 Environment and Engineering: Conda, Docker, dependency planning, shell execution, Git workflows, Overleaf, harness utilities, and skill updates.
1.3 Research Discovery: academic literature search, multi-engine evidence collection, research idea generation, and method design support.
2.1 Structural and Functional Toolchains: FreeSurfer, FSL, Nilearn, fMRIPrep, CONN, and HCP Pipelines for reconstruction, segmentation, registration, GLM, ICA, and connectivity.
2.2 Diffusion and Electrophysiology: DIPY, QSIPrep, and MNE support denoising, tensor metrics, tractography, EEG cleaning, and feature extraction.
2.3 Visualization and Clinical Interop: brain network visualization, zALFF regional summaries, FreeSurfer mesh export, and DICOM-compatible output conversion.
3.1 sMRI: T1/T2 preprocessing, tissue segmentation, cortical parcellation, FreeSurfer surfaces, and WMH segmentation from FLAIR+T1w inputs.
3.2 fMRI: preprocessing, denoising, ROI time series, seed/ROI connectivity, task GLM, resting-state ICA, and effective connectivity routes.
3.3 DWI + EEG: eddy correction, diffusion tensor metrics, tractography, connectome construction, EEG artifact removal, spectral analysis, and feature extraction.
4.1 ADNI: acquisition guidance, BIDS organization, preprocessing, ROI analysis, phenotype modeling, and derived dataset generation.
4.2 HCP: structural, functional, and diffusion execution across HCP Pipelines with multimodal result integration.
4.3 UK Biobank and Expansion Targets: UKB post-extraction brain analysis plus routes that can extend toward ADHD200, ABCD, ABIDE, COBRE, PNC, HBN, and UCLA datasets.
5.1 Deep Learning: BrainGNN, FM-APP, and NeuroStorm for phenotype prediction and neuroimaging representation learning.
5.2 Statistical and Classical ML: GLM, ICA, DictLearning, SVM, SpaceNet, K-means, hierarchical clustering, filtering, and detrending.
5.3 Model Routing: unified model entry with preprocessing dependency checks, input/output mapping, executable plans, and benchmark-friendly evaluation steps.
6.1 End-to-End Pipelines: single-modality and dataset-specific chains for sMRI, fMRI, DWI, EEG, ADNI, HCP, and UKB analysis.
6.2 Experiment Control: METHOD-to-experiment execution, Git-based setup, ablations, checkpoints, verification, audit logs, and resumable runs.
6.3 Research Output: manuscript writing, clean dialogue logs, visualization assets, reproducibility reports, and maintainable skill-library updates.
Use NeuroClaw to prototype auditable neuroimaging pipelines and autonomous experiment loops in your lab.